Our laboratory aims to unravel the mechanisms that tumor cells initially employ to escape the control of their environment and initiate malignant growth, as well as those they utilize later to survive drug treatments.
We are pursuing these mechanisms using single-cell transcriptomics, an approach that offers an unbiased way to study them at multiple levels: in individual cells, subpopulations, and tissues. Single-cell transcriptomics also enables us to analyze aberrations, whether readily detectable in cancer specimens or generated through genome editing, and their effects on transcriptional networks that define cellular states, on a scale sufficient to yield both biologically and clinically relevant information.
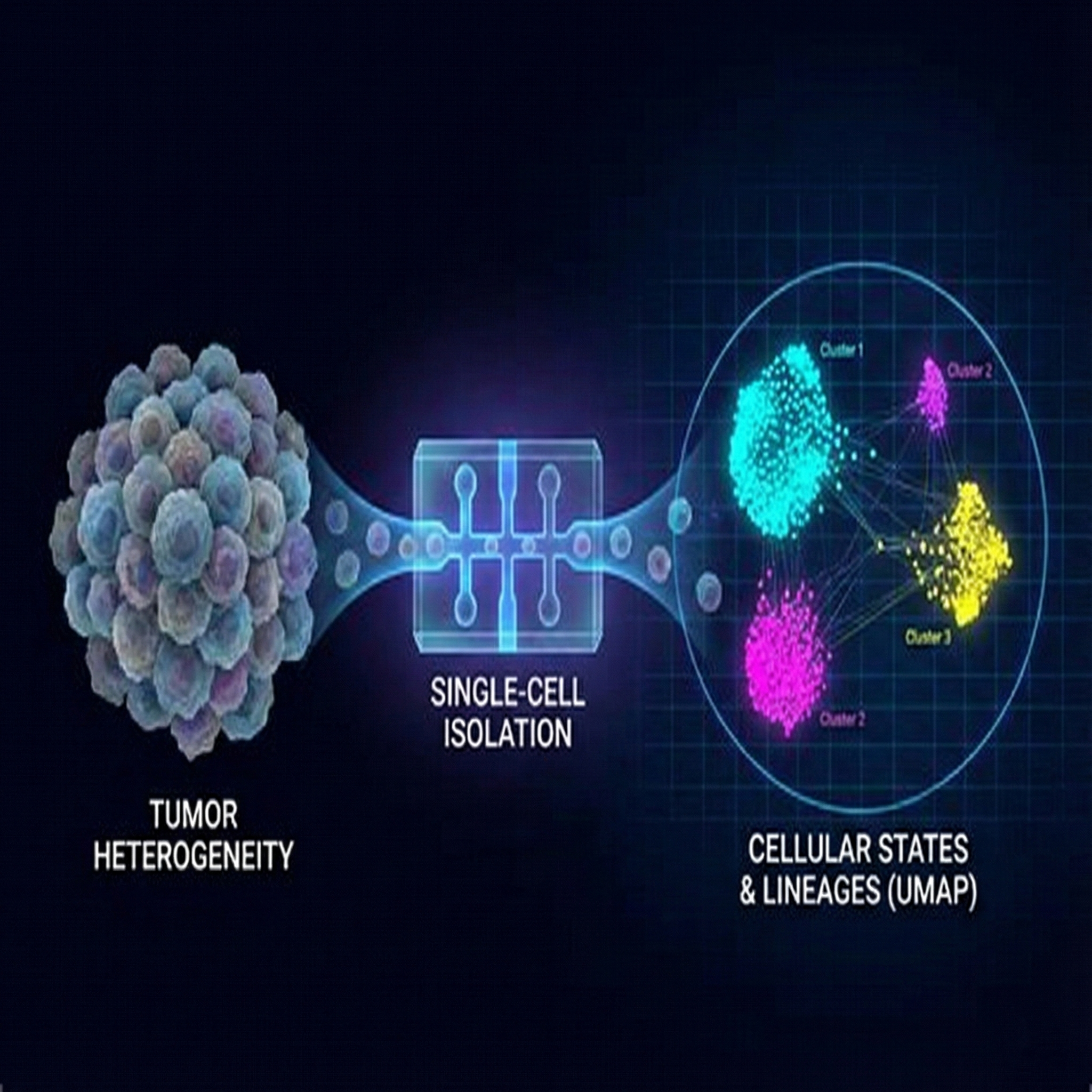
Single-cell technologies to decode tumor heterogeneity
Integrating scRNA-seq, lineage tracing, and computational modeling to map cellular states and predict treatment responses.
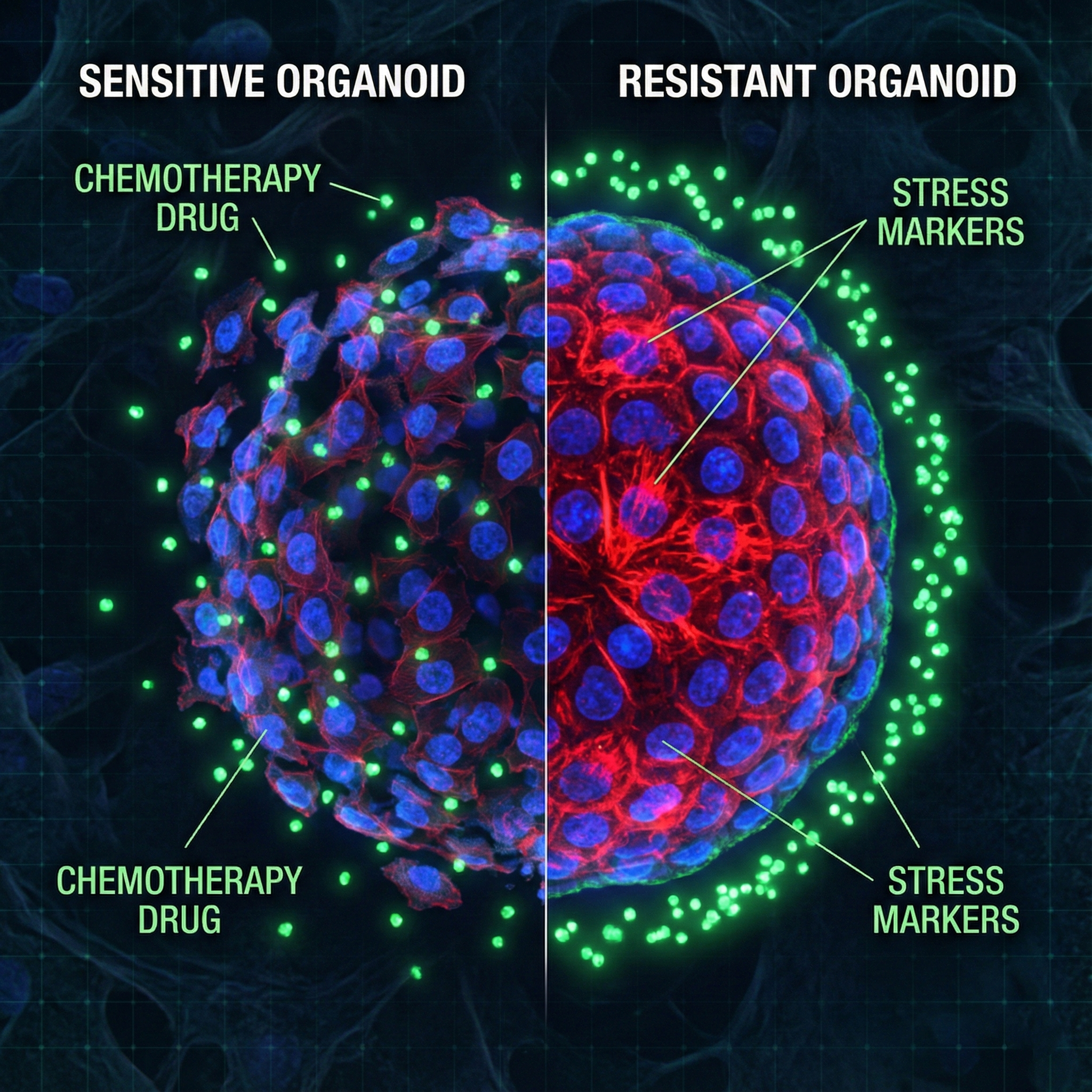
Mechanisms of chemotherapy resistance in ovarian cancer
Dissecting intrinsic, primed, and adaptive resistance using organoid models, stress-recording tools, and genetic perturbations.

Computational frameworks for cancer biology
Developing machine-learning and statistical models to infer cell fate, transcription factor activity, and responses to perturbations.

Spatial organization of the tumor microenvironment
Combining spatial imaging and transcriptomics to reveal cell–cell interactions that shape tumor evolution.
Current Projects
What doesn’t kill you: Overcoming primed and adaptive resistance in ovarian cancer (STRONGER), 2024-2029
This ERC-funded project by Anna Vähärautio studies how past stress encounters induce treatment resistance in ovarian cancer cells. The project will reveal biographies of individual cancer cells to understand how they become resilient. How do previous adaptations to various stressors shape the adaptation during anti-cancer treatment? Is it possible to block treatment-induced adaptation by correctly timed co-treatments.
Sister cell lineage tracing to predict pre-existing resistance
We have several sub-projects that apply ReSisTrace, our lineage tracing method that takes advantage of the similarity of sister cells to couple cell state to its future fate. These projects are funded by the Cancer Foundation and Academy of Finland (collaboration with Jing Tang, Markus Vähä-Koskela and Hanna Seppälä also involving pancreatic cancer organoids).
Single-cell characterization of high-grade serous ovarian cancer
Using the scRNA-seq data from clinical HGSOC tumor specimens we have accumulated so far, we study how various features of the tumors affect their drug response. We are especially interested in the immune and non-immune tumor microenvironment, cancer cell states in relation to those of normal post-menopausal fimbria (the tissue of origin for HGSOC), as well as how distinct genetic drivers modulate HGSOC cell states and their treatment responses.